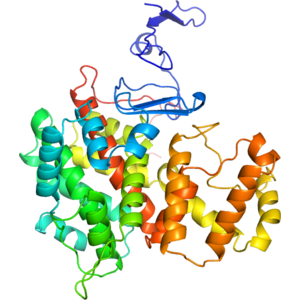
Structure of IDP91458
Type II citrate synthase from Vibrio vulnificus.
Annotation
- Description
- Citrate synthases catalyze the first reaction in the citric acid cycle (the Krebs' cycle), namely the condensation of acetyl-coenzyme A with oxaloacetate to form citrate and coenzyme A. This reaction is important for energy generation and for carbon assimilation. The enzymes are found in two distinct structural types: type I enzymes (found in eukaryotes, Gram-positive bacteria and Archaea) forming homodimers and type II enzymes, which are found in Gram-negative bacteria and are hexameric in structure. The enzyme monomer is composed of two alpha-helical domains. The cleft between these domains forms the active site. Type II enzymes possess an extra N-terminal beta-sheet domain.
- Functional assignment
- transferase
Ligands
| Ligand code | Name | Ligand type |
|---|---|---|
| MSE | modified residue | |
| 175 | 3,5-dihydro-5-methylidene-4h-imidazol-4-on |
Structure information
Unit cell parameters
- Space Group
- P 63 2 2
- Unit Cell
-
a=109.72Å, b=109.72Å, c=153.65Å
α=90.00, β=90.00, γ=120.00 - Solvent content
- Matthews coefficient
- Resolution range
- 95.02-2.50Å (2.56-2.50Å)
- Rall(%)
- 19.1
- Rwork(%)
- 18.9 (25.3)
- Rfree(%)
- 23.1 (34.4)
- Num. observed reflections
- 19512 (1239)
- Num. Rfree reflections
- 995 (60)
- Completeness(%)
- 99.6 (99.6)
- Num Atoms
- 3433
- Num Waters
- 50
- Num Hetatoms
- 0
- Model mean isotropic B factor
- 52.480Å2
- RMSD bond length
- 0.014Å
- RMSD bond angle
- 1.673°
- Filename uploaded
- idp91458.pdb (uploaded on Mar 16, 2012 12:01 PM)
- Inserted
- Mar 16, 2012