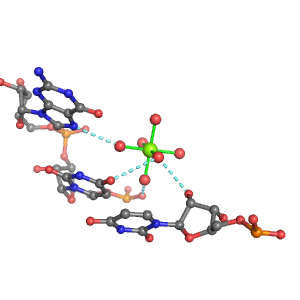
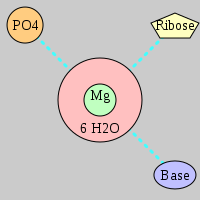
| Mg2+ ion |
Site type |
Inner-sphere ligands |
Outer-sphere ligands |
|---|
| PDB ID |
Mg2+ ID |
Oph |
Or |
Ob+Nb |
Water |
Other |
Pout |
Rout |
Bout |
|
2ao5 |
A198 |
Pout·Rout·Bout |
|
|
|
A228, A249, A250, A251 |
|
G:A6 |
G:A6 |
G:A6 |
|
2qbh |
A1565 |
Pout·Rout·Bout |
|
|
|
A1707, A1708, A1709, A1710, A1711, A1712 |
|
G:A727 |
C:A853 |
G:A727 |
|
2qou |
A1566 |
Pout·Rout·Bout |
|
|
|
A2011, A2012, A2013, A2014, A2015, A2016 |
|
A:A1357 |
A:A1254 |
C:A1359 |
|
2qou |
A1579 |
Pout·Rout·Bout |
|
|
|
A2074, A2075, A2076, A2077, A2078, A2079 |
|
U:A916 |
U:A13 |
G:A917 |
|
2qp0 |
A1592 |
Pout·Rout·Bout |
|
|
|
A2122, A2123, A2124, A2125, A2126, A2127 |
|
A:A1357 |
A:A1254 |
C:A1359 |
|
3v26 |
A1731 |
Pout·Rout·Bout |
|
|
|
A2035, A2036, A2037, A2038, A2039, A2040 |
|
A:A915 |
U:A13 |
G:A916 |
|
3v2d |
A3608 |
Pout·Rout·Bout |
|
|
|
A5412, A5413, A5414, A5415 |
|
A:A1762 |
G:A1681 |
U:A1757 |
|
4bpe |
A5066 |
Pout·Rout·Bout |
|
|
|
A2425, A2426, A2429, A2449, A2450, A2471 |
|
U:A1529 |
G:A1531 |
G:A1531 |
|
4bpe |
A5020 |
Pout·Rout·Bout |
|
|
|
A2034, A2035, A2036, A2058, A2059, A2269 |
|
A:A101 |
A:A100 |
U:A297 |
|
4bpe |
A5026 |
Pout·Rout·Bout |
|
|
|
A2037, A2057, A2067, A2278, A2279, A2280 |
|
A:A104 |
G:A374 |
U:A294 |
|
4bpn |
A5026 |
Pout·Rout·Bout |
|
|
|
A6439, A6440, A6441, A6442, A6443, A6444 |
|
A:A104 |
G:A374 |
U:A294 |
|
4bpn |
A5066 |
Pout·Rout·Bout |
|
|
|
A6528, A6529, A6530, A6531, A6532, A6534 |
|
U:A1529 |
G:A1531 |
G:A1531 |
|
4bpn |
A5077 |
Pout·Rout·Bout |
|
|
|
A6606, A6607, A6608, A6609, A6610, A6611 |
|
U:A1275 |
U:A1277 |
U:A1277 |
|
4bpn |
A5056 |
Pout·Rout·Bout |
|
|
|
A6201, A6202, A6203, A6204, A6205, A6206 |
|
G:A1113 |
U:A8 |
G:A1113 |
|
4bpn |
A5020 |
Pout·Rout·Bout |
|
|
|
A6401, A6403, A6404, A6405, A6406, A6407 |
|
A:A101 |
A:A100 |
U:A297 |
|
4bpo |
A5066 |
Pout·Rout·Bout |
|
|
|
A2285, A2286, A2287, A2444, A2445, A2446 |
|
U:A1529 |
G:A1531 |
G:A1531 |
|
4bpo |
A5020 |
Pout·Rout·Bout |
|
|
|
A2035, A2036, A2037, A2057, A2356, A2357 |
|
A:A101 |
A:A100 |
U:A297 |
|
4bpo |
A5026 |
Pout·Rout·Bout |
|
|
|
A2038, A2039, A2056, A2065, A2365, A2366 |
|
A:A104 |
G:A374 |
U:A294 |
|
4bpp |
A5020 |
Pout·Rout·Bout |
|
|
|
A7331, A7332, A7333, A7334, A7335, A7336 |
|
A:A101 |
A:A100 |
U:A297 |
|
4bpp |
A5026 |
Pout·Rout·Bout |
|
|
|
A7427, A7428, A7429, A7430, A7431, A7432 |
|
A:A104 |
G:A374 |
U:A294 |
|
4bpp |
A5066 |
Pout·Rout·Bout |
|
|
|
A8067, A8068, A8069, A8070, A8071, A8072 |
|
U:A1529 |
G:A1531 |
G:A1531 |
|
4bpp |
A5056 |
Pout·Rout·Bout |
|
|
|
A7907, A7908, A7909, A7910, A7911, A7912 |
|
G:A1113 |
U:A8 |
G:A1113 |
|
4dr2 |
A1747 |
Pout·Rout·Bout |
|
|
|
A2372, A2373, A2374, A2375, A2376 |
|
G:A666 |
A:A665 |
G:A667 |
|
4dr3 |
A1719 |
Pout·Rout·Bout |
|
|
|
A2111, A2112, A2113, A2114 |
|
A:A1004 |
U:A1025 |
G:A1024 |
|
4dv3 |
A1851 |
Pout·Rout·Bout |
|
|
|
A2229, A2230, A2231, A2232, A2233, A2234 |
|
G:A1224 |
C:A1223 |
A:A977 |
|