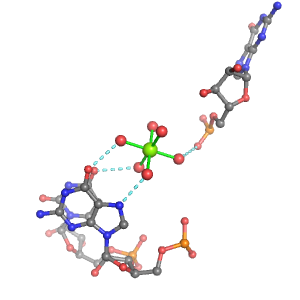
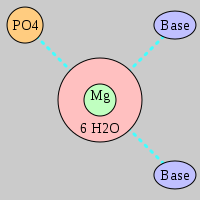
| Mg2+ ion |
Site type |
Inner-sphere ligands |
Outer-sphere ligands |
|---|
| PDB ID |
Mg2+ ID |
Oph |
Or |
Ob+Nb |
Water |
Other |
Pout |
Rout |
Bout |
|
1egk |
C3 |
Pout·2Bout |
|
|
|
B61, C28, C29, C30, C31, C32 |
|
DA:B45 |
|
A:C104, G:C105 |
|
1ehz |
A570 |
Pout·2Bout |
|
|
|
A731, A732, A733, A734, A735, A736 |
|
C:A25 |
|
U:A41, G:A42 |
|
1evv |
A904 |
Pout·2Bout |
|
|
|
A936, A937, A938, A939, A940, A941 |
|
C:A25 |
|
G:A42, U:A41 |
|
1vs5 |
A1557 |
Pout·2Bout |
|
|
|
A1666, A1667, A1668, A1669, A1670, A1671 |
|
A:A1046 |
|
G:A1047, G:A1048 |
|
1vs5 |
A1564 |
Pout·2Bout |
|
|
|
A1702, A1703, A1704, A1705, A1706, A1707 |
|
A:A781 |
|
G:A799, G:A800 |
|
1vs6 |
B2997 |
Pout·2Bout |
|
|
|
B3438, B3439, B3440, B3441, B3442, B3443 |
|
U:B686 |
|
G:B770, G:B771 |
|
1vs7 |
A1562 |
Pout·2Bout |
|
|
|
A1689, A1690, A1691, A1692, A1693, A1694 |
|
G:A1079 |
|
G:A1077, U:A1076 |
|
1vs7 |
A1564 |
Pout·2Bout |
|
|
|
A1701, A1702, A1703, A1704, A1705, A1706 |
|
A:A315 |
|
G:A319, G:A318 |
|
1vt2 |
A3002 |
Pout·2Bout |
|
|
|
A3503, A3504, A3505, A3506, A3507, A3508 |
|
C:A2616 |
|
G:A2049, C:A2047 |
|
1y26 |
X101 |
Pout·2Bout |
|
|
|
X267, X268, X269, X270, X271, X272 |
|
A:X45 |
|
G:X46, U:X47 |
|
1yls |
B244 |
Pout·2Bout |
|
|
|
B367, B368, B369, B370, B371, B372 |
|
U:B212 |
|
G:B213, U:B231 |
|
2avy |
A1562 |
Pout·2Bout |
|
|
|
A1693, A1694, A1695, A1696, A1697, A1698 |
|
G:A266 |
|
G:A257, G:A258 |
|
2avy |
A1556 |
Pout·2Bout |
|
|
|
A1665, A1666, A1667, A1668, A1669, A1670 |
|
A:A1046 |
|
G:A1047, G:A1048 |
|
2aw7 |
A1563 |
Pout·2Bout |
|
|
|
A1702, A1703, A1704, A1705, A1706, A1707 |
|
A:A315 |
|
G:A319, G:A318 |
|
2aw7 |
A1561 |
Pout·2Bout |
|
|
|
A1690, A1691, A1692, A1693, A1694, A1695 |
|
G:A1079 |
|
G:A1077, U:A1076 |
|
2i2p |
A1545 |
Pout·2Bout |
|
|
|
A1614, A1615, A1616, A1617, A1618, A1619 |
|
G:A803 |
|
G:A771, U:A772 |
|
2i2t |
B3018 |
Pout·2Bout |
|
|
|
B3531, B3532, B3533, B3534, B3535, B3536 |
|
G:B2597 |
|
G:B2595, U:B1971 |
|
2i2t |
B3019 |
Pout·2Bout |
|
|
|
B3538, B3539, B3540, 4754, 4756, 4759 |
|
A:B2740 |
|
G:B2742, U:B2743 |
|
2i2v |
B2957 |
Pout·2Bout |
|
|
|
B3276, B3277, B3278, B3279, B3280, B3281 |
|
A:B2386 |
|
C:B2326, A:B2386 |
|
2i2v |
B2917 |
Pout·2Bout |
|
|
|
B3085, B3086, B3087, B3088, B3089, B3090 |
|
A:B2033 |
|
G:B2027, U:B2028 |
|
2i2v |
B2969 |
Pout·2Bout |
|
|
|
B3337, B3338, B3339, B3340, B3341, B3342 |
|
G:B649 |
|
C:B650, G:B651 |
|
2i2v |
B3018 |
Pout·2Bout |
|
|
|
B3530, B3531, B3532, B3533, B3534, B3535 |
|
G:B2597 |
|
G:B2595, U:B1971 |
|
2otj |
08100 |
Pout·2Bout |
|
|
|
03788, 04283, 05543, 06186, 07847 |
|
C:02077 |
|
G:02079, G:02080 |
|
2qal |
A2077 |
Pout·2Bout |
|
|
|
A2078, A2079, A2080, A2081, A2082, A2083 |
|
A:A1046 |
|
G:A1047, G:A1048 |
|
2qan |
A2134 |
Pout·2Bout |
|
|
|
A2135, A2136, A2137, A2138, A2139, A2140 |
|
A:A1046 |
|
G:A1047, G:A1048 |
|