| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
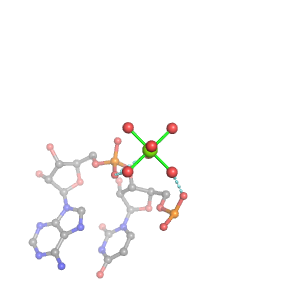
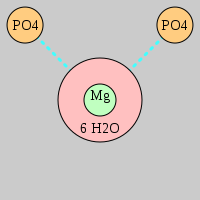
| Site type used in the classification is a string abbreviation of ligand composition in Mg2+ coordination sphere | |||
| Abbreviations used for Mg2+ "Site type" |
Mg2+ inner-sphere ligands | Geometry for inner-sphere OP | Mg2+ outer-sphere moieties |
|---|---|---|---|
|
Oph phosphate oxygen (OP1/OP2) Or ribose oxygen (O2'/O4') or oxygen bridging phosphate and ribose (O3'/O5') Ob nucleobase oxygen Nb nucleobase nitrogen |
cis- two OP ligands adopt cis- isoform trans- two OP ligands adopt trans- isoform fac- three OP ligands adopt fac- isoform mer- three OP ligands adopt mer- isoform |
Pout phosphate moiety Rout ribose moiety Bout nucleobase moiety | |
| Reference: | Zheng H, Shabalin IG, Handing KB, Bujnicki JM, Minor W. (2015) Magnesium binding architectures in RNA crystal structures: validation, binding preferences, classification, and motif detection. Nucleic Acid Research 43(7):3789-801 [Pubmed]. | ||
Developed by Heping Zheng and Ivan Shabalin at Minor lab (http://olenka.med.virginia.edu/CrystUVa/)
For any help please contact <dust@iwonka.med.virginia.edu>