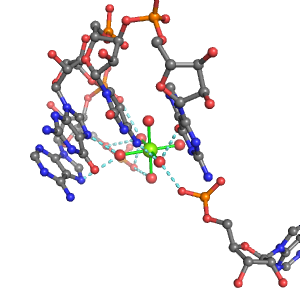
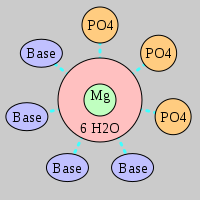
| Mg2+ ion |
Site type |
Inner-sphere ligands |
Outer-sphere ligands |
|---|
| PDB ID |
Mg2+ ID |
Oph |
Or |
Ob+Nb |
Water |
Other |
Pout |
Rout |
Bout |
|
1vs8 |
B2938 |
3Pout·4Bout |
|
|
|
B3180, B3181, B3182, B3183, B3184, B3185 |
|
A:B447, A:B28, U:B29 |
|
G:B473, U:B29, C:B31, G:B474 |
|
2qam |
B3110 |
3Pout·4Bout |
|
|
|
B3700, B3701, B3702, B3703, B3704, B3705 |
|
A:B28, A:B447, U:B29 |
|
U:B29, G:B473, G:B30, G:B474 |
|
2qao |
B3320 |
3Pout·4Bout |
|
|
|
B3881, B3882, B3883, B3884, B3885, B3886 |
|
A:B582, G:B583, A:B1253 |
|
A:B582, G:B583, C:B584, G:B585 |
|
2qbe |
B3110 |
3Pout·4Bout |
|
|
|
B3701, B3702, B3703, B3704, B3705, B3706 |
|
A:B28, A:B447, U:B29 |
|
U:B29, G:B473, G:B474, G:B30 |
|
2qbi |
B2983 |
3Pout·4Bout |
|
|
|
B3384, B3385, B3386, B3387, B3388, B3389 |
|
A:B2451, C:B2452, A:B2453 |
|
A:B2453, G:B2454, A:B2497, G:B2455 |
|
2qbi |
B2923 |
3Pout·4Bout |
|
|
|
B3102, B3103, B3104, B3105, B3106, B3107 |
|
U:B29, A:B28, A:B447 |
|
U:B29, G:B473, G:B30, G:B474 |
|
2qov |
B2983 |
3Pout·4Bout |
|
|
|
B3388, B3389, B3390, B3391, B3392, B3393 |
|
C:B2452, A:B2453, A:B2451 |
|
A:B2453, G:B2454, A:B2497, G:B2455 |
|
2qov |
B2956 |
3Pout·4Bout |
|
|
|
B3261, B3262, B3263, B3264, B3265, B3266 |
|
A:B582, A:B1253, G:B583 |
|
A:B582, C:B584, G:B585, G:B583 |
|
2qoz |
B2923 |
3Pout·4Bout |
|
|
|
B3597, B3598, B3599, B3600, B3601, B3602 |
|
U:B29, A:B28, A:B447 |
|
G:B473, G:B474, U:B29, G:B30 |
|
2qoz |
B2983 |
3Pout·4Bout |
|
|
|
B3878, B3879, B3880, B3881, B3882, B3883 |
|
A:B2451, C:B2452, A:B2453 |
|
A:B2497, G:B2454, G:B2455, A:B2453 |
|
2z4l |
B2984 |
3Pout·4Bout |
|
|
|
B3386, B3387, B3388, B3389, B3390, B3391 |
|
A:B2451, C:B2452, A:B2453 |
|
A:B2497, G:B2454, G:B2455, A:B2453 |
|
2z4n |
B2931 |
3Pout·4Bout |
|
|
|
B3141, B3142, B3143, B3144, B3145, B3146 |
|
A:B2451, A:B2453, C:B2452 |
|
A:B2453, G:B2454, A:B2497, G:B2455 |
|
3ofz |
A2905 |
3Pout·4Bout |
|
|
|
A3044, A3045, A3046, A3047, A3048, A3049 |
|
U:A29, A:A447, A:A28 |
|
G:A473, G:A474, U:A29, G:A30 |
|
3r8t |
A2905 |
3Pout·4Bout |
|
|
|
A3076, A3077, A3078, A3079, A3080, A3081 |
|
U:A29, A:A28, A:A447 |
|
G:A474, U:A29, G:A473, G:A30 |
|
3v23 |
A3608 |
3Pout·4Bout |
|
|
|
A5042, A5043, A5044, A5045, A5046, A5047 |
|
U:A29, A:A447, A:A28 |
|
G:A474, G:A473, U:A29, G:A30 |
|
3v2d |
A3402 |
3Pout·4Bout |
|
|
|
A4838, A4839, A4840, A4841, A4842, A4843 |
|
U:A29, A:A447, A:A28 |
|
G:A474, G:A473, U:A29, G:A30 |
|
4a17 |
0325 |
3Pout·4Bout |
|
|
|
08371, 08372, 08373, 08374, 08375, 08376 |
|
G:365, G:366, A:364 |
|
U:3109, G:3108, G:365, G:366 |
|
4gar |
A3002 |
3Pout·4Bout |
|
|
|
A3207, A3208, A3209, A3210, A3211, A3212 |
|
U:A29, A:A447, A:A28 |
|
U:A29, G:A30, C:A31, G:A474 |
|
4tp1 |
A3072 |
3Pout·4Bout |
|
|
|
A3527, A3528, A3529, A3530, A3531, A3532 |
|
C:A2452, A:A2453, A:A2451 |
|
A:A2453, G:A2454, G:A2455, A:A2497 |
|
4tp3 |
A3080 |
3Pout·4Bout |
|
|
|
A3557, A3558, A3559, A3560 |
|
A:A2407, A:A2406, U:A2408 |
|
G:A2410, A:A2411, U:A2408, G:A2409 |
|
4tp7 |
A3081 |
3Pout·4Bout |
|
|
|
A3557, A3558, A3559, A3560 |
|
A:A2406, G:A411, A:A2407 |
|
A:A2411, A:A2407, U:A2408, G:A2409 |
|
4tpb |
A3080 |
3Pout·4Bout |
|
|
|
A3557, A3558, A3559, A3560 |
|
G:A411, A:A2407, A:A2406 |
|
G:A2410, A:A2411, U:A2408, G:A2409 |
|
4tpf |
A3072 |
3Pout·4Bout |
|
|
|
A3522, A3523, A3524, A3525, A3526, A3527 |
|
A:A2453, A:A2451, C:A2452 |
|
A:A2497, A:A2453, G:A2454, G:A2455 |
|
4tpf |
A3003 |
3Pout·4Bout |
|
|
|
A3207, A3208, A3209, A3210, A3211, A3212 |
|
U:A29, A:A28, A:A447 |
|
U:A29, G:A473, G:A30, C:A31 |
|