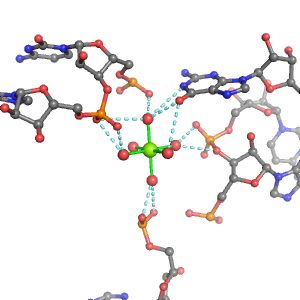
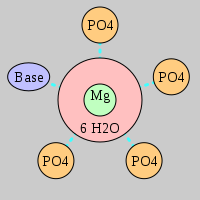
| Mg2+ ion |
Site type |
Inner-sphere ligands |
Outer-sphere ligands |
|---|
| PDB ID |
Mg2+ ID |
Oph |
Or |
Ob+Nb |
Water |
Other |
Pout |
Rout |
Bout |
|
1vs8 |
B2909 |
4Pout·Bout |
|
|
|
B3035, B3036, B3037, B3038, B3039, B3040 |
|
A:B1937, G:B1959, A:B1936, C:B1958 |
|
G:B1959 |
|
1vs8 |
B2910 |
4Pout·Bout |
|
|
|
B3041, B3042, B3043, B3044, B3045, B3046 |
|
C:B1986, A:B1669, A:B1987, C:B2551 |
|
G:B1954 |
|
1vt2 |
A2956 |
4Pout·Bout |
|
|
|
A3281, A3282, A3283, A3284, A3285, A3286 |
|
A:A1669, C:A1986, C:A2551, A:A1987 |
|
G:A1954 |
|
2aw4 |
B2970 |
4Pout·Bout |
|
|
|
B3322, B3323, B3324, B3325, B3326, B3327 |
|
A:B1669, C:B1986, A:B1987, C:B2551 |
|
G:B1954 |
|
2awb |
B2909 |
4Pout·Bout |
|
|
|
B3036, B3037, B3038, B3039, B3040, B3041 |
|
A:B1937, G:B1959, A:B1936, C:B1958 |
|
G:B1959 |
|
2i2p |
A1559 |
4Pout·Bout |
|
|
|
A1681, A1682, A1683, A1684, A1685, A1686 |
|
U:A13, A:A16, U:A14, G:A15 |
|
U:A14 |
|
2i2t |
B2968 |
4Pout·Bout |
|
|
|
B3321, B3322, B3323, B3324, B3325, B3326 |
|
C:B1351, U:B1352, G:B1377, C:B1376 |
|
G:B1377 |
|
2i2t |
B2970 |
4Pout·Bout |
|
|
|
B3332, B3333, B3334, B3335, B3336, B3337 |
|
A:B1669, C:B1986, C:B2551, A:B1987 |
|
G:B1954 |
|
2i2t |
B2931 |
4Pout·Bout |
|
|
|
B3150, B3151, B3152, B3153, B3154, B3155 |
|
A:B1937, C:B1958, G:B1959, A:B1936 |
|
G:B1959 |
|
2i2u |
A1556 |
4Pout·Bout |
|
|
|
A1662, A1663, A1664, A1665, A1666, A1667 |
|
A:A768, U:A804, C:A805, A:A767 |
|
G:A769 |
|
2i2v |
B2910 |
4Pout·Bout |
|
|
|
B3051, B3052, B3053, B3054, B3055, B3056 |
|
C:B1986, A:B1669, A:B1987, C:B2551 |
|
G:B1954 |
|
2i2v |
B2920 |
4Pout·Bout |
|
|
|
B3100, B3101, B3102, B3103, B3104, B3105 |
|
C:B1351, U:B1352, G:B1377, C:B1376 |
|
G:B1377 |
|
2qam |
B3181 |
4Pout·Bout |
|
|
|
B3758, B3759, B3760, B3761, B3762, B3763 |
|
C:B2050, A:B2051, G:B2578, A:B2614 |
|
A:B2614 |
|
2qao |
B3033 |
4Pout·Bout |
|
|
|
B3645, B3646, B3647, B3648, B3649, B3650 |
|
C:B2551, A:B1987, A:B1669, C:B1986 |
|
G:B1954 |
|
2qba |
B3382 |
4Pout·Bout |
|
|
|
B3920, B3921, B3922, B3923, B3924, B3925 |
|
C:B1986, A:B1669, C:B2551, A:B1987 |
|
G:B1954 |
|
2qbc |
B3033 |
4Pout·Bout |
|
|
|
B3646, B3647, B3648, B3649, B3650, B3651 |
|
A:B1669, C:B1986, C:B2551, A:B1987 |
|
G:B1954 |
|
2qbc |
B3344 |
4Pout·Bout |
|
|
|
B3899, B3900, B3901, B3902, B3903, B3904 |
|
A:B1916, A:B1912, A:B1913, U:B1915 |
|
A:B1918 |
|
2qbc |
B3485 |
4Pout·Bout |
|
|
|
B4014, B4015, B4016, B4017, B4018, B4019 |
|
C:B2050, A:B2614, A:B2051, G:B2578 |
|
A:B2614 |
|
2qbg |
B2910 |
4Pout·Bout |
|
|
|
B3043, B3044, B3045, B3046, B3047, B3048 |
|
A:B1669, C:B1986, C:B2551, A:B1987 |
|
G:B1954 |
|
2qbi |
B2970 |
4Pout·Bout |
|
|
|
B3322, B3323, B3324, B3325, B3326, B3327 |
|
A:B1669, C:B1986, A:B1987, C:B2551 |
|
G:B1954 |
|
2qbi |
B2935 |
4Pout·Bout |
|
|
|
B3160, B3161, B3162, B3163, B3164, B3165 |
|
C:B2050, G:B2578, A:B2051, A:B2614 |
|
A:B2051 |
|
2qbi |
B2968 |
4Pout·Bout |
|
|
|
B3311, B3312, B3313, B3314, B3315, B3316 |
|
C:B1376, C:B1351, U:B1352, G:B1377 |
|
G:B1377 |
|
2qbk |
B2963 |
4Pout·Bout |
|
|
|
B3295, B3296, B3297, B3298, B3299, B3300 |
|
U:B1915, A:B1916, A:B1912, A:B1913 |
|
A:B1918 |
|
2qov |
B2935 |
4Pout·Bout |
|
|
|
B3161, B3162, B3163, B3164, B3165, B3166 |
|
G:B2578, C:B2050, A:B2051, U:B2613 |
|
A:B2614 |
|
2qp1 |
B2963 |
4Pout·Bout |
|
|
|
B3803, B3804, B3805, B3806, B3807, B3808 |
|
U:B1915, A:B1916, A:B1912, A:B1913 |
|
U:B1917 |
|